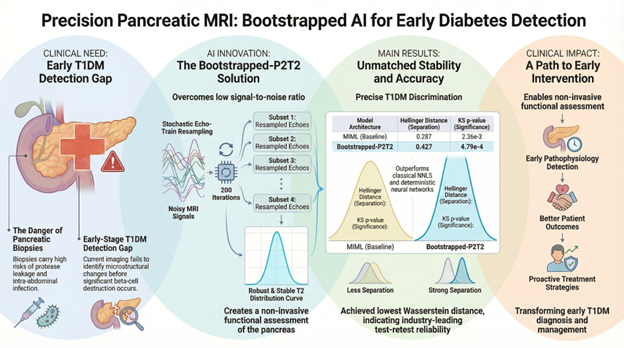
AI for Quantitative Pancreatic MRI in Diabetes: Robust T2 relaxometry using physics-informed deep learning
PI-Technion, Assoc. Prof. Moti Freiman PI-Rambam, Prof. Ram Weiss, MD
A Technion–Rambam collaboration advancing non-invasive imaging biomarkers for diabetes through a novel AI framework that enables stable and sensitive analysis of pancreatic microstructure from MRI data. Why It Matters Type 1 diabetes develops years before clinical diagnosis, yet no imaging modality today can detect early pancreatic changes such as inflammation or beta-cell loss.
Current MRI approaches
- Struggle with low signal-to-noise ratio (SNR) in abdominal imaging
- Fail to capture subtle microstructural tissue changes
- Provide limited ability to generate reliable quantitative biomarkers
This creates a critical barrier to early detection, monitoring, and treatment personalization in diabetes.
Our AI Approach
We introduce a bootstrapped, physically-primed deep learning framework for multi-component T2 relaxometry in MRI.
Core innovation:
- Physics-informed neural networks integrating MRI acquisition parameters
- Inference-time bootstrapping to reduce noise and improve robustness
- Estimation of full T2 distributions rather than single scalar values
- Transformation of deterministic models into probabilistic ensemble predictors
This approach enables stable, high-fidelity characterization of pancreatic tissue microstructure, even under low-SNR conditions.
Research Goals
- Develop motion-robust, quantitative MRI biomarkers for pancreatic tissue
- Enable early detection of diabetes-related microstructural changes
- Improve reproducibility and sensitivity of MRI-based measurements
- Establish a framework for clinical translation of qMRI in metabolic disease
Key Results
- Achieved significantly improved test–retest stability compared to classical and deep-learning baselines
- Demonstrated the strongest statistical separation between T1DM patients and healthy controls
- Reduced sensitivity to noise through bootstrap-based inference, outperforming standard ensemble methods
- Enabled detection of physiologically meaningful changes following glucose intake
Notably, the proposed method achieved the lowest statistical p-values and highest distributional separation, indicating superior biomarker sensitivity.
Impact
This work establishes a new paradigm for quantitative MRI in metabolic disease, enabling:
- Non-invasive detection of early pancreatic dysfunction
- Development of imaging biomarkers for diabetes progression
- Extension to oncology and other abdominal imaging applications
Publication
Ben Atya H. et al., Bootstrapped Physically-Primed Neural Networks for Robust T2 Distribution Estimation in Low-SNR Pancreatic MRI, MIDL 2026 (Full Paper), open access version: https://arxiv.org/abs/2603.14084
Acknowledgment
This research was supported in part by the Technion EVPR Foundation for Collaborative Research between the Technion and the Rambam Health Care Campus.
MRI data were acquired at the May-Blum-Dah Human MRI research center at the Technion